Density, distribution function, quantile function and random generation for a univariate distribution estimated by maximum likelihood.
Usage
dml(x, obj, log = FALSE)
pml(q = q, obj, lower.tail = TRUE, log.p = FALSE)
qml(p = p, obj, lower.tail = TRUE, log.p = FALSE)
rml(n = n, obj)Arguments
- x, q
vector of quantiles.
- obj
an
univariateMLobject.- log, log.p
logical; if
TRUE, the probabilities p are gives aslog(p).- lower.tail
logical; if
TRUE(default), the probabilities are \(P[X \le x]\) otherwise, \(P[X > x]\)- p
vector of probabilities.
- n
number of observations. If
length(n) > 1, the length is taken to be the number required.
Value
dml gives the density, pml gives the distribution
function, qml gives the quantile function, and rml
generates random deviates.
Details
dml is the density, pml is the distribution function,
qml is the quantile function, and rml is the random variable
generator.
These functions work like their counterparts in stats, e.g.
Normal. The univariateML object contains both maximum
likelihood estimates and the identity of the model these estimates were
calculated under. These functions are wrappers around underlying density,
distribution, quantile and random generation functions where unknown
parameters are filled with the maximum likelihood estimates.
See the example.
Examples
## Simple example
obj <- mlnorm(airquality$Wind)
#> Loading required package: intervals
dml(0.5, obj) == dnorm(0.5, mean = obj[1], sd = obj[2])
#> [1] TRUE
obj <- mlbetapr(airquality$Wind)
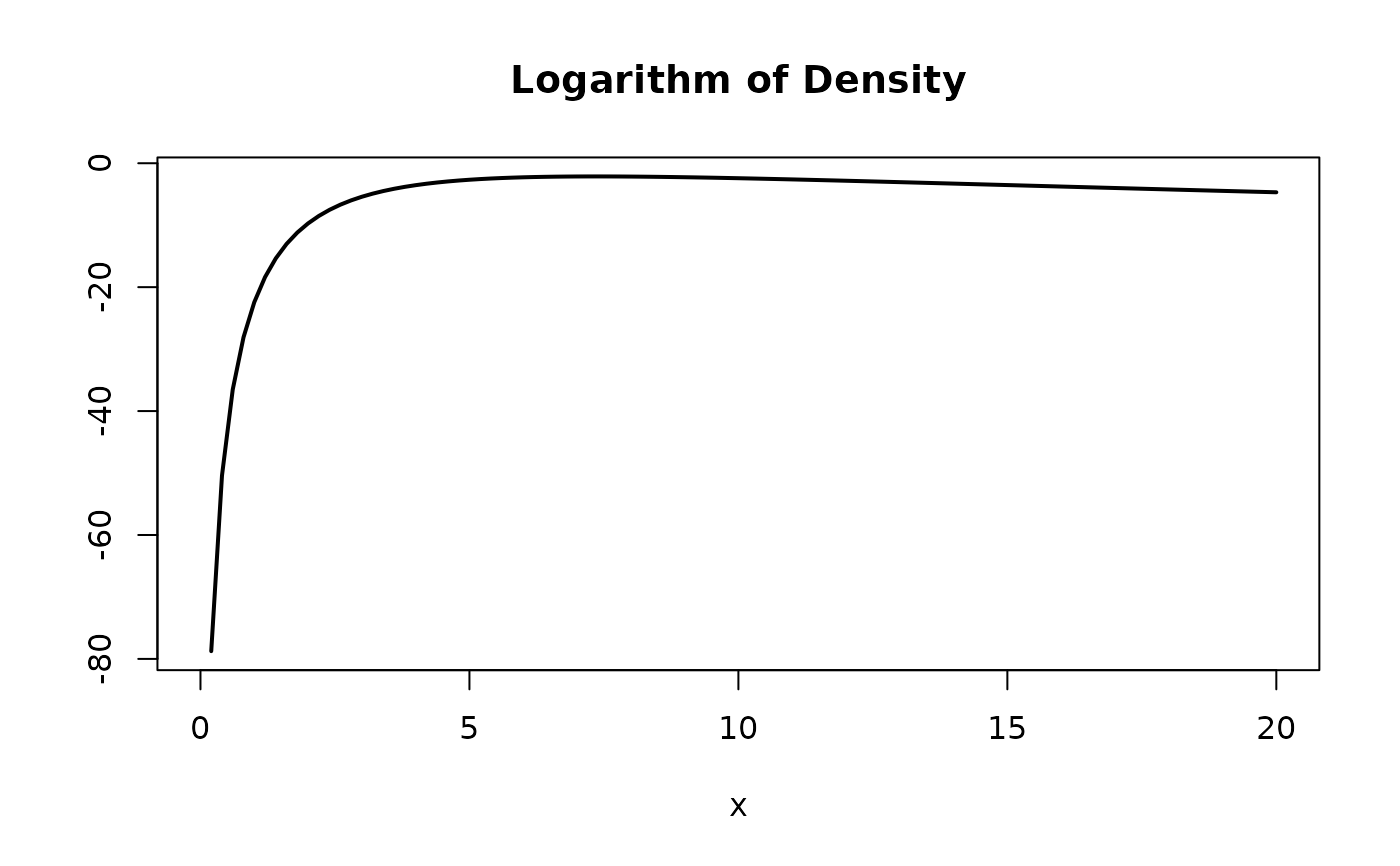
# Plot the logarithm of the beta prime distribution.
plot(\(x) dml(x, obj, log = TRUE),
from = 0, to = 20,
main = "Logarithm of Density", ylab = NA, lwd = 2
)